Does Life Evolve Like a Great Branching Tree?
By Helen Wong 王思齊

There is grandeur in this view of life … from so simple a beginning endless forms most beautiful and most wonderful have been, and are being, evolved.”
Charles Darwin, On the Origin of Species [1]
Darwin’s theory of evolution by natural selection explains how life evolves through inherited changes passed from one generation to the next. When it comes to inheritance, we often think of genes being passed down vertically from parents to offspring. This pattern, known as “vertical gene transfer,” has long been considered the main way genetic information moves through populations. However, it is not the only way: Genes can also move horizontally between unrelated organisms in a process called “horizontal gene transfer” (HGT). Once thought to be rare, HGT is now known to occur across nearly all branches of life and plays a key role in how life adapts in fast and sometimes surprising ways [2–3].
HGT in Prokaryotes
While the term HGT might be new to you, chances are you have already heard of one of its consequences: antibiotic resistance in pathogenic bacteria. This is one of the most famous examples in action.
Bacteria (and possibly archaea) can acquire “foreign” antibiotic resistance genes and other genetic materials through three main processes: transformation, transduction, and conjugation (figure 1). In transformation, bacteria take up naked DNA from their surroundings, often in the form of plasmids. Transduction involves the transfer of genes between bacterial hosts by bacteriophages, which are viruses that infect bacteria. DNA can also be transferred from one bacterium to another through conjugation, during which bacteria mate by forming a cell-to-cell connection. HGT is so widespread among bacteria that researchers estimate that around one-third of all gene transfers in bacterial evolution happen this way [4].
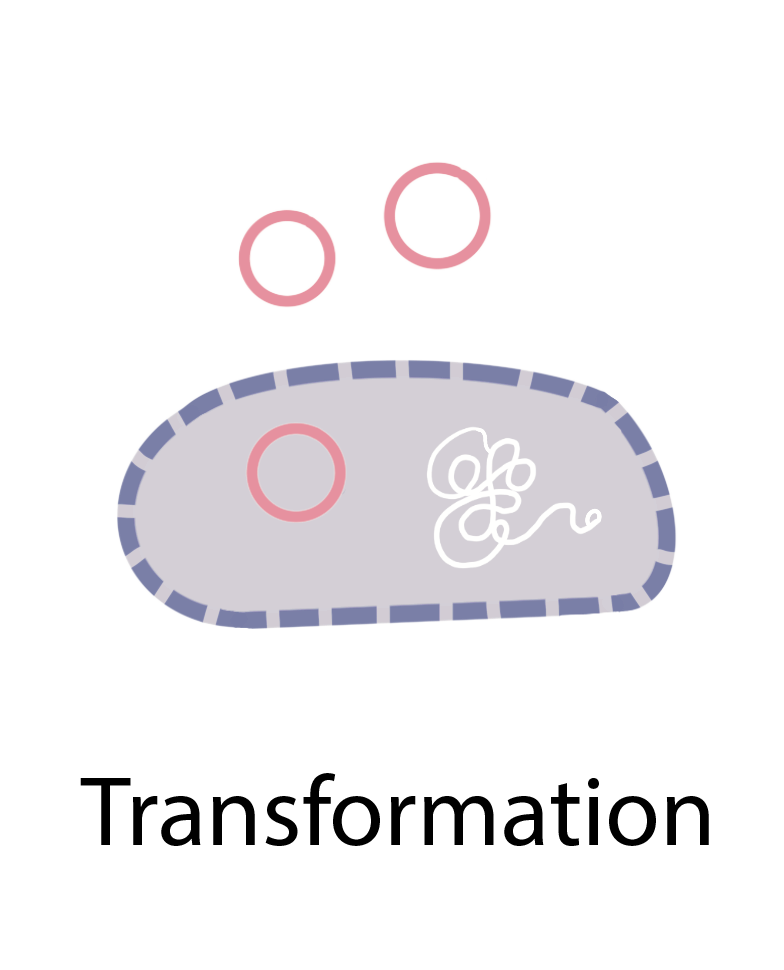 | 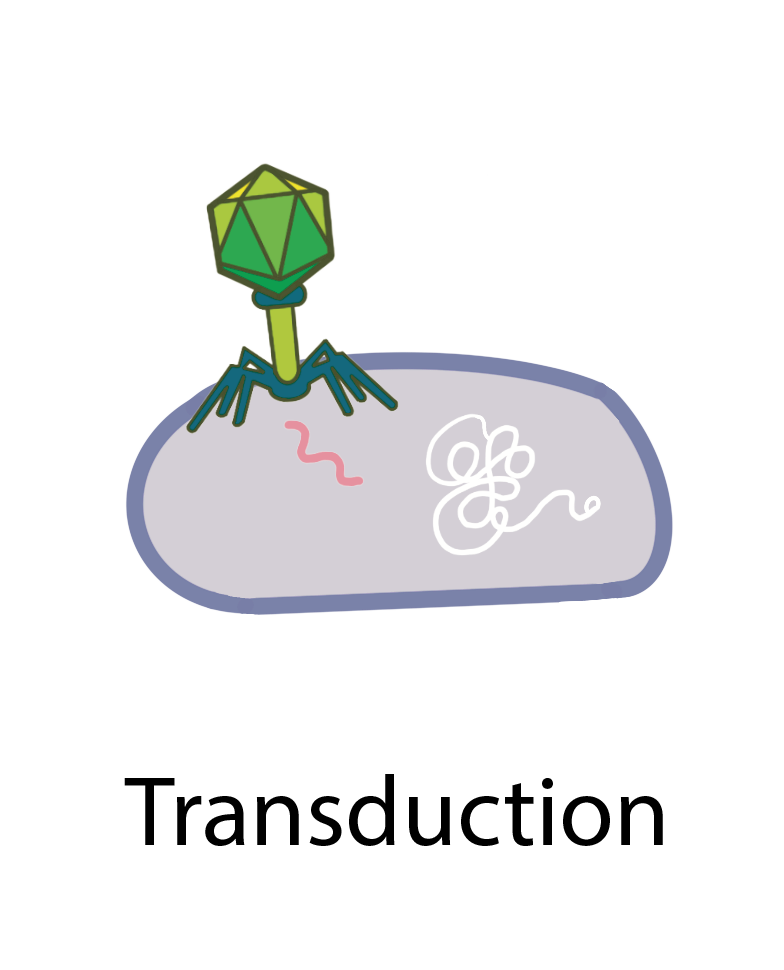 | 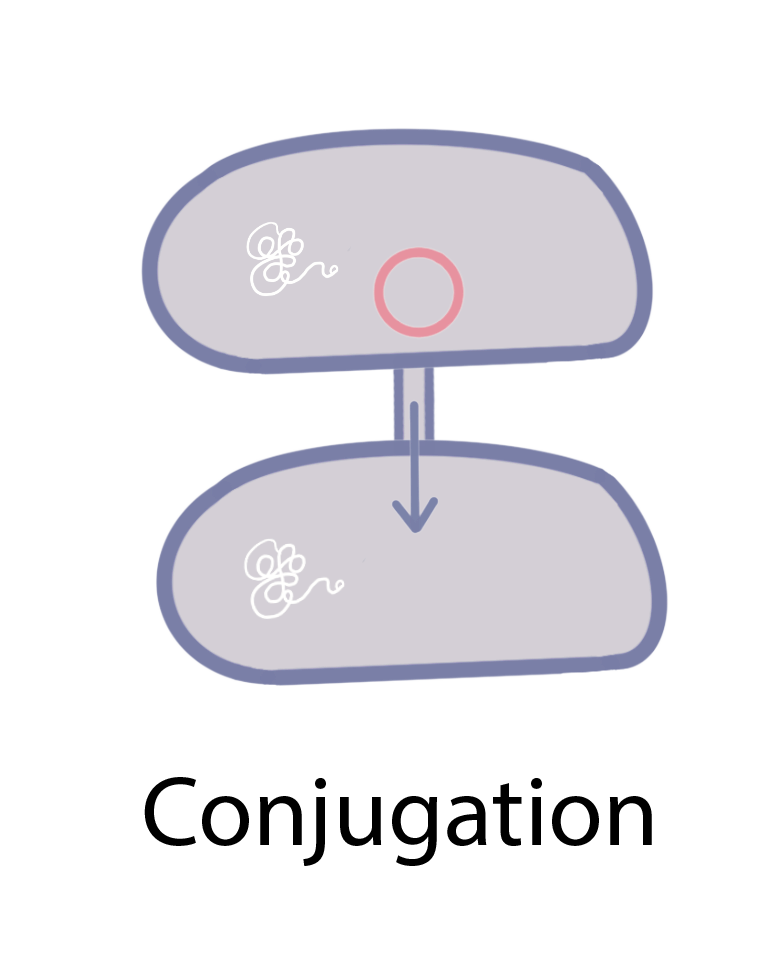 |
Figure 1 Illustration of the three major mechanisms of horizontal gene transfer in prokaryotes: transformation, transduction, and conjugation.
HGT in Eukaryotes
However, things are different for eukaryotes, such as animals, plants, and fungi. For most eukaryotes, germ cells are kept separate from somatic cells [2–3]. This means that if a foreign gene ends up in a somatic cell, it won’t be passed down to the next generation; only genes that incorporate into sperms and eggs, or their progenitor cells (cells that will divide into sperms and eggs), can be inherited. Therefore, for HGT to have lasting effects in eukaryotes, it often has to happen in a germ cell.
Still, HGT can occur in eukaryotes, and certain conditions seem to increase its likelihood. These include processes like phagocytosis, where cells engulf other cells or particles, and symbiosis, where two different organisms live in close association. HGT may also happen more frequently during early developmental stages when the organism only consists of one or a few unspecialized cells, such as when it is a spore, zygote, or embryo.
Unlike bacteria, the main way foreign DNA becomes part of a eukaryotic genome is through a process called “non-homologous end joining.” This is a natural DNA repair mechanism that fixes breaks by directly connecting the broken ends of DNA. When foreign DNA is present during this repair process, it can sometimes be inserted into the genome.
Although biologists haven’t reached a consensus on how common HGT is in eukaryotes, some fascinating examples have been documented. One involves an aphid ancestor acquiring carotenoid genes from fungi [5]. This was the first time a group of carotenoid genes was found in an animal genome. These pigment-producing genes allow aphids to produce red, green, and yellow colors. In nature, red and green aphids are both common because predators and parasites tend to prefer one color over the other, thus retaining the color variation.
A more recent example of HGT in eukaryotes involves a newly discovered system similar to CRISPR-Cas systems [6–7]. CRISPR is a naturally occurring DNA sequence found in prokaryotic genomes, derived from fragments of past invading viral DNA [8]. During subsequent infections, a family of DNA-cutting enzymes (or more technically “endonucleases”) called Cas uses complementary RNA transcribed from CRISPR sequences as a guide to recognize the invader’s DNA, and then destroy matching viral DNA to protect themselves. While researchers wondered whether such RNA-guided systems only exist in prokaryotes, they found the eukaryotic counterpart in fungi. At its core is an RNA-guided endonuclease called Fanzor. Interestingly, scientists discovered that the Fanzor genes may have originated from prokaryotic tnpB genes, which were likely transferred to eukaryotes through HGT.
Detecting HGT Events
So, how do scientists actually find these examples of HGT? The key lies in looking closely at the gene that was horizontally transferred. Since it originally came from another species, it should still look similar to the version in the original donor species in terms of the amino acid sequence encoded.
This idea is the basis for a common method called “phylogenetic conflict [2–3].” In simple terms, scientists compare two kinds of “family trees”: one for the species themselves, and one for the gene in question. If a gene was transferred from one species to another, the gene tree might show two species as closely related, even if the species tree says otherwise. For example, in figure 2, if gene X was transferred from species C to species D, the gene X tree would wrongly suggest that C and D are close relatives as they share a more similar gene X.

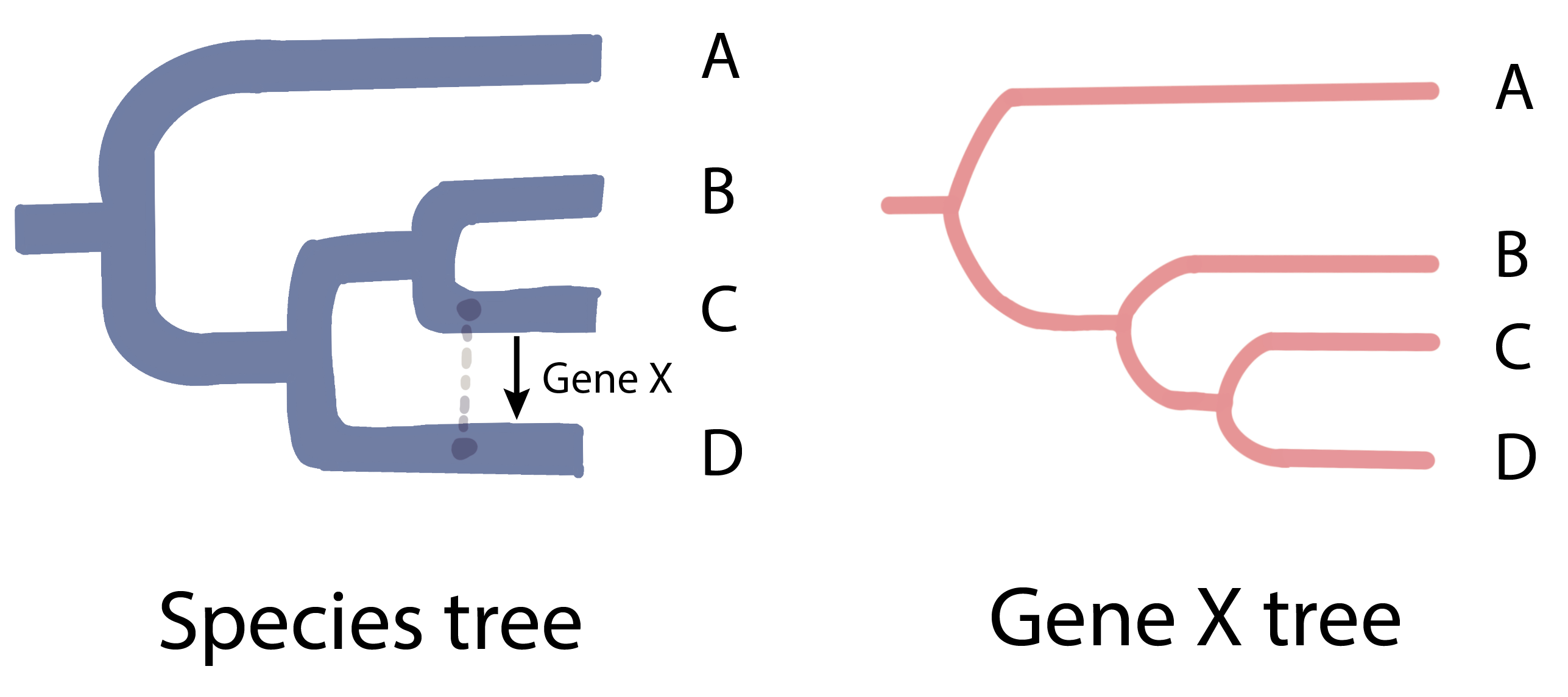
Figure 2 Phylogenetic conflict arising from a mismatch between a species tree and a gene tree. If gene X was horizontally transferred from species C to species D, the gene X tree would wrongly suggest that C and D are close relatives as they share a more similar gene X.
But a tree mismatch isn’t always enough to prove that HGT happened. There could be other reasons for the odd pattern, such as contamination during DNA sequencing [3]. To be more certain, scientists would check where gene X is located in the genome of species D. If gene X is found right next to other genes that are already known to belong to species D, that’s a good sign it’s truly a part of D’s genome, rather than just a stray piece of DNA from somewhere else. It also helps to see if gene X has common features of a real gene, like introns (non-coding sequences commonly found in eukaryotic genes). On top of that, RNA sequencing data can show if the gene is actually expressed and used by the cell.
HGT and the Tree of Life
The phylogenetic conflicts introduced by HGT have led some biologists to question whether Darwin’s tree of life model still fully captures the complexity of evolutionary relationships, especially among prokaryotes [9]. In Darwin’s metaphor, life evolves like a great branching tree, with species diverging from common ancestors over time. HGT adds a new dimension to this picture: It causes some branches to reach sideways, connecting distant twigs in unexpected ways. This lateral exchange creates a more web-like structure, leading some to propose a network model of evolution rather than a strictly branching tree. Yet whether life is ultimately represented as a tree, a network, or a mix of both, one thing remains clear: HGT has played a profound role in shaping the evolution of the “endless forms most beautiful and most wonderful.”
References
[1] Darwin, C. (1859). On the origin of species by means of natural selection, or, The preservation of favoured races in the struggle for life (1st ed.). John Murray.
[2] Zhaxybayeva, O., & Doolittle, W. F. (2011). Lateral gene transfer. Current Biology, 21(7), R242–R246. https://doi.org/10.1016/j.cub.2011.01.045
[3] Husnik, F., & McCutcheon, J. P. (2018). Functional horizontal gene transfer from bacteria to eukaryotes. Nature Reviews Microbiology, 16(2), 67–79. https://doi.org/10.1038/nrmicro.2017.137
[4] Coleman, G. A., Davín, A. A., Mahendrarajah, T. A., Szánthó, L. L., Spang, A., Hugenholtz, P., Szöllősi, G. J., & Williams, T. A. (2021). A rooted phylogeny resolves early bacterial evolution. Science, 372(6542). https://doi.org/10.1126/science.abe0511
[5] Moran, N. A., & Jarvik, T. (2010). Lateral transfer of genes from fungi underlies carotenoid production in aphids. Science, 328(5978), 624–627. https://doi.org/10.1126/science.1187113
[6] Saito, M., Xu, P., Faure, G., Maguire, S., Kannan, S., Altae-Tran, H., Vo, S., Desimone, A., Macrae, R. K., & Zhang, F. (2023). Fanzor is a eukaryotic programmable RNA-guided endonuclease. Nature, 620(7974), 660–668. https://doi.org/10.1038/s41586-023-06356-2
[7] Eisenstadt, L. (2023, June 28). Researchers uncover a new CRISPR-like system in animals that can edit the human genome. MIT News. https://news.mit.edu/2023/fanzor-system-in-animals-can-edit-human-genome-0628
[8] Broad Institute. (n.d.). Questions and answers about CRISPR. https://www.broadinstitute.org/what-broad/areas-focus/project-spotlight/questions-and-answers-about-crispr
[9] Blais, C., & Archibald, J. M. (2021). The past, present and future of the tree of life. Current Biology, 31(7), R314–R321. https://doi.org/10.1016/j.cub.2021.02.052